RNASeqR: RNA-Sequencing pipeline R package development
Introduction
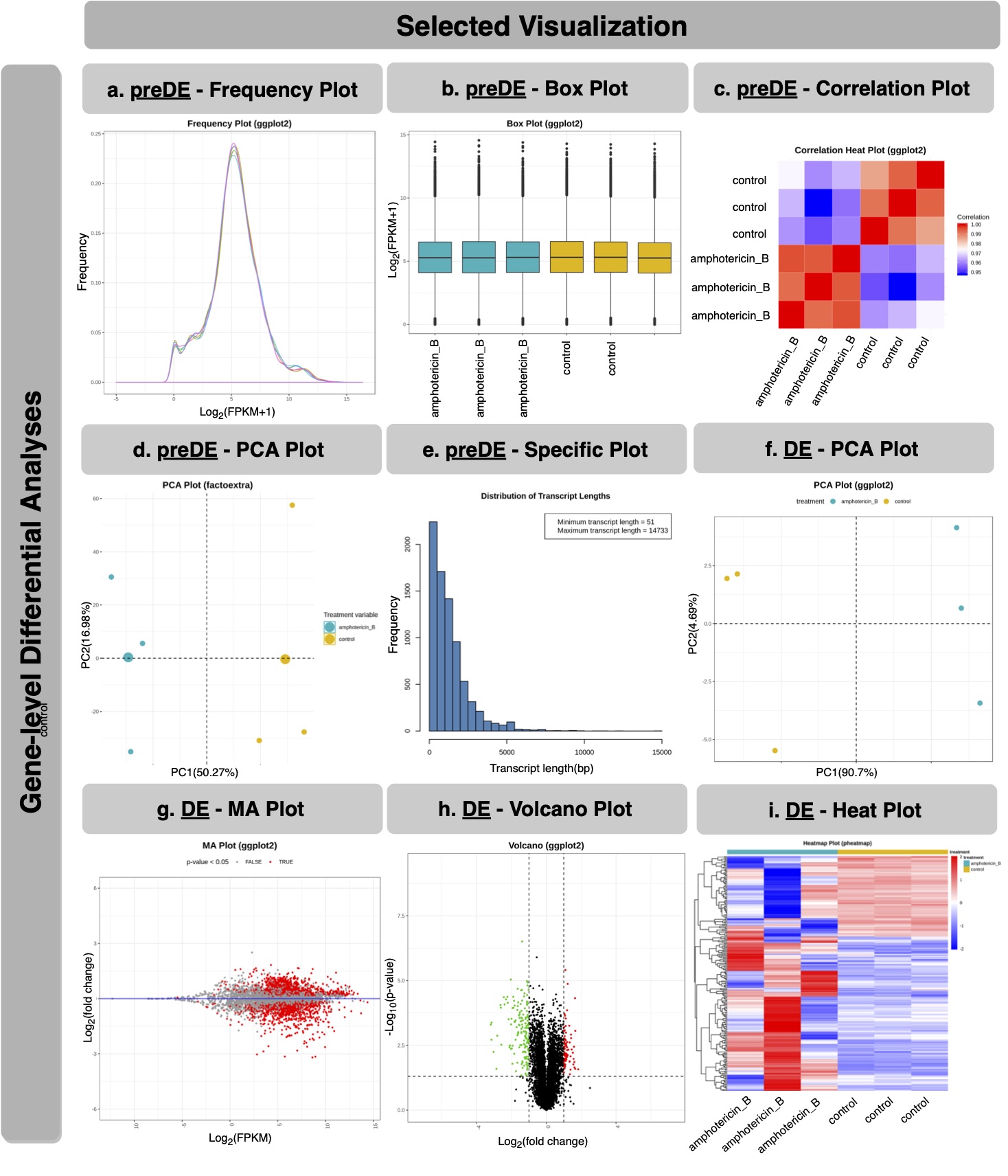
I developed RNASeqR for two-group (case-control) RNA-Seq analysis and it is now available on Bioconductor 3.10 release. There are six steps: “RNASeqRParam S4 Object Creation”, “Environment Setup”, “Quality Assessment”, “Reads Alignment & Quantification”, “Gene-level Differential Analyses” and “Functional Analyses”. Each step corresponds to a function in this package. After running functions in order, a basic RNASeq analysis would be done easily.
In December, 2018, I presented this work in NTU Centers of Genomics and Precision Medicine (NTUCGM) Summer Research competition and won the first prize among all summer research students. In June, 2019, I oral presented this work in ICIBM 2019, Columbus, USA. The manuscript is now published in IEEE/ACM Transactions on Computational Biology and Bioinformatics.
Source code & Documentation
RNASeqR: an R package for automated two-group RNA-Seq analysis workflow
Kuan-Hao Chao, Y.W. Hsiao, Y.F. Lee, C.Y. Lee, L.C. Lai, M.H. Tsai, T.P. Lu, and E.Y. Chuang* (2019). RNASeqR: an R package for automated two-group RNA-Seq analysis workflow, IEEE/ACM Transactions on Computational Biology and Bioinformatics, vol. 18, no. 5, pp. 2023-2031, 1 Sept.-Oct. 2021, https://10.1109/TCBB.2019.2956708
[ICIBM 2019] International Conference on Intelligent Biology and Medicine 2019
15-min presentation at International Conference on Intelligent Biology and Medicine (ICIBM 2019), Columbus, Ohio
[NTUCGM 2018] Summer Research Competition 2018
30-min presentation at NTU Centers of Genomics and Precision Medicine (NTUCGM) Summer Research, Taipei, Taiwan
